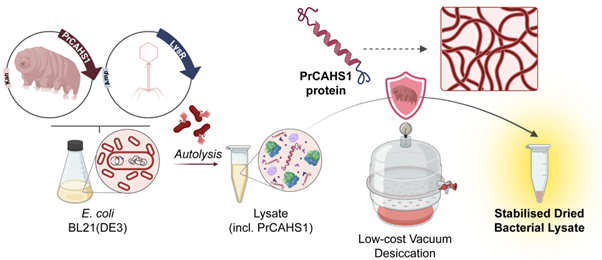
Tardigrade-Derived Strategy for Low-Cost Storage of Cell-Free Expression Lysates.
Meckelburg M*, Banlaki I*, Gaizauskaite A, Niederholtmeyer H#
bioRxiv preprint. (2026)
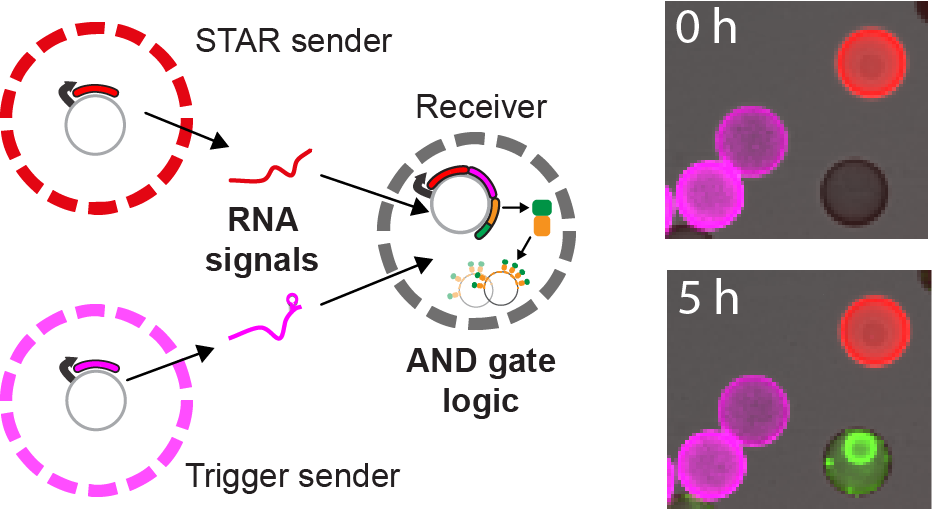
RNA-based communication in heterogeneous populations of cell mimics.
Lehr FX, Banlaki I, Bumüller E, Mercier T, Niederholtmeyer H#
ACS Synthetic Biology, 15, 2, 574–585. (2026)
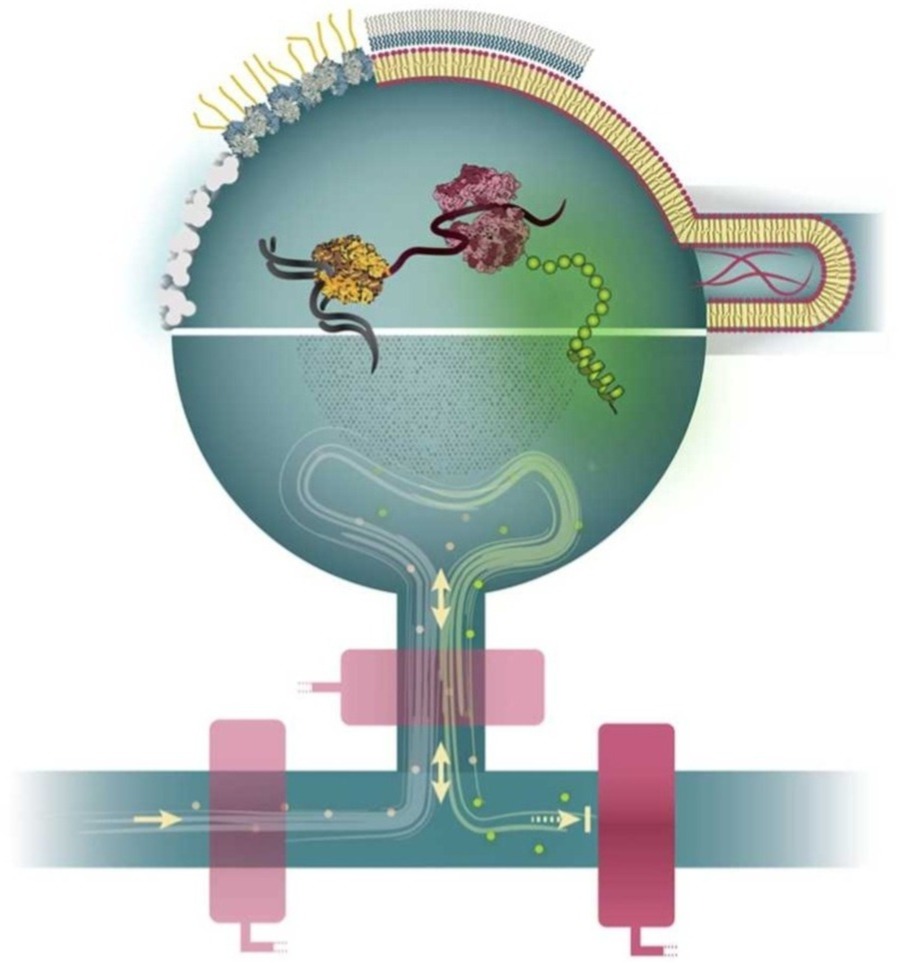
Controlled protein synthesis and spatial organisation in microfluidic environments.
Gaizauskaite A, Crean EE, Banlaki I, Kalkowski JL, Niederholtmeyer H#
Current Opinion in Biotechnology. (2025)
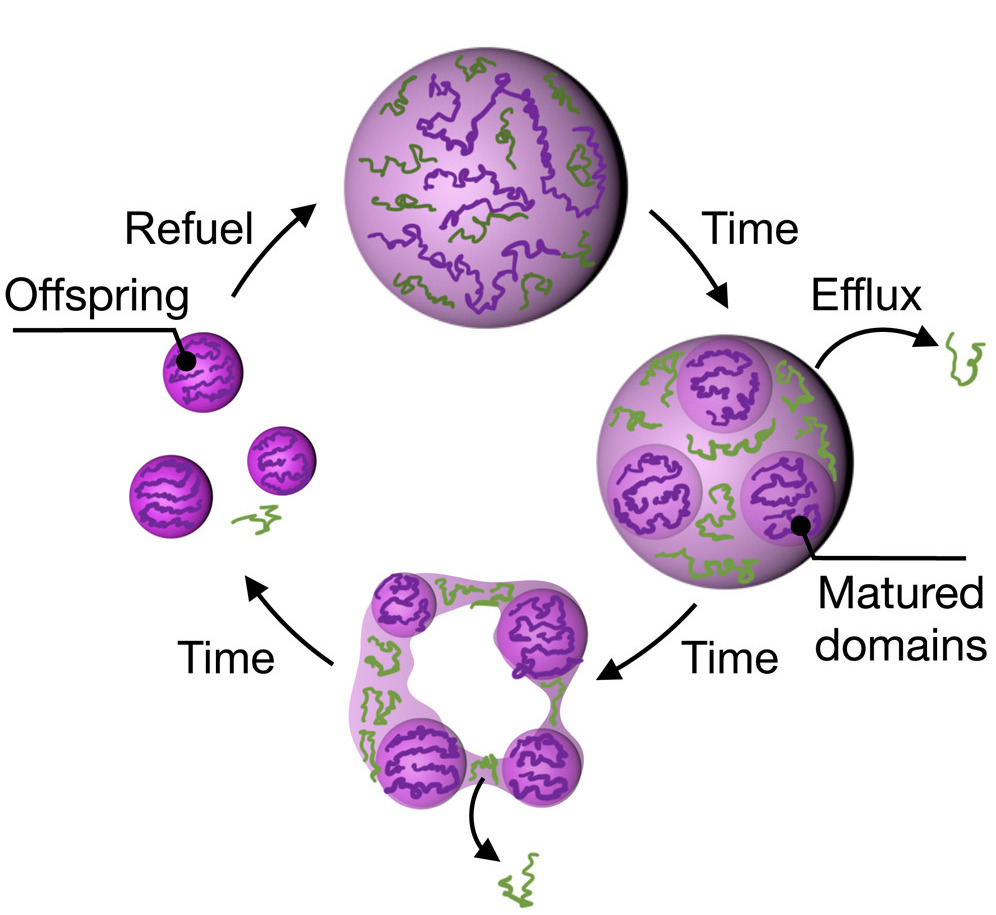
Toward synthetic life—Emergence, growth, creation of offspring, decay, and rescue of fuel-dependent synthetic cells.
Wenisch M, Li Y, Braun MG, Eylert L, Späth F, Poprawa SM, Rieger B, Synatschke CV, Niederholtmeyer H, Boekhoven J#.
Chem. (2025)
![]()
A programmable biomimetic cytoskeleton.
Niederholtmeyer H#
Nature Chemistry 17, 311–313. (2025)
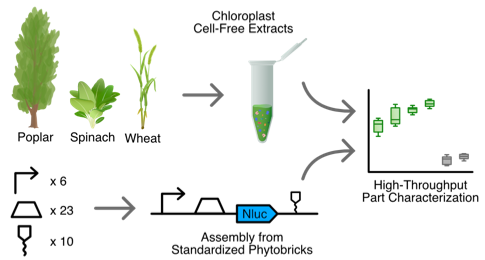
Chloroplast cell-free systems from different plant species as a rapid prototyping platform.
Böhm CV*, Inckemann R*#, Burgis M, Baumann J, Brinkmann CK, Lipińska KE, Gilles S, Freudigmann J, Seiler V, Clark LG, Jewett MC, Voll LM#, Niederholtmeyer H#
ACS Synthetic Biology 13, 2412-2424. (2024)
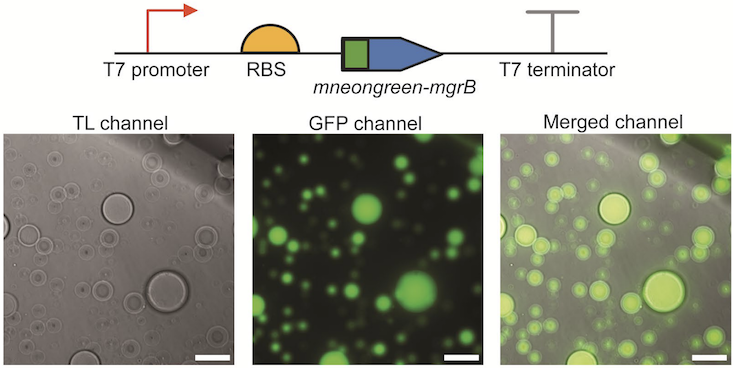
A cell-free system for functional studies of small membrane proteins.
Jiang S, Çelen G, Glatter T, Niederholtmeyer H#, Yuan J#
Journal of Biological Chemistry, 300 (11). (2024)
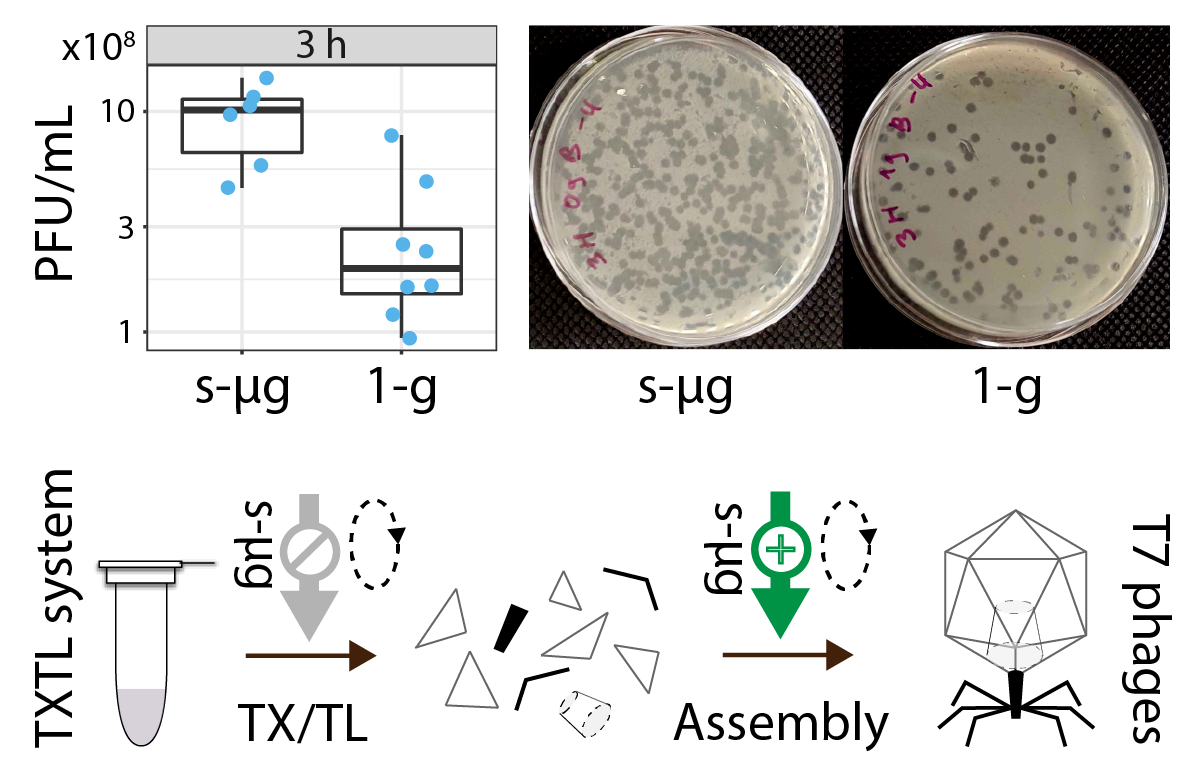
Enhanced assembly of bacteriophage T7 produced in cell-free reactions under simulated microgravity.
Lehr FX*, Pavletic B*, Glatter T, Heimerl T, Moeller R#, Niederholtmeyer H#
npj Microgravity 10, 30. (2024)
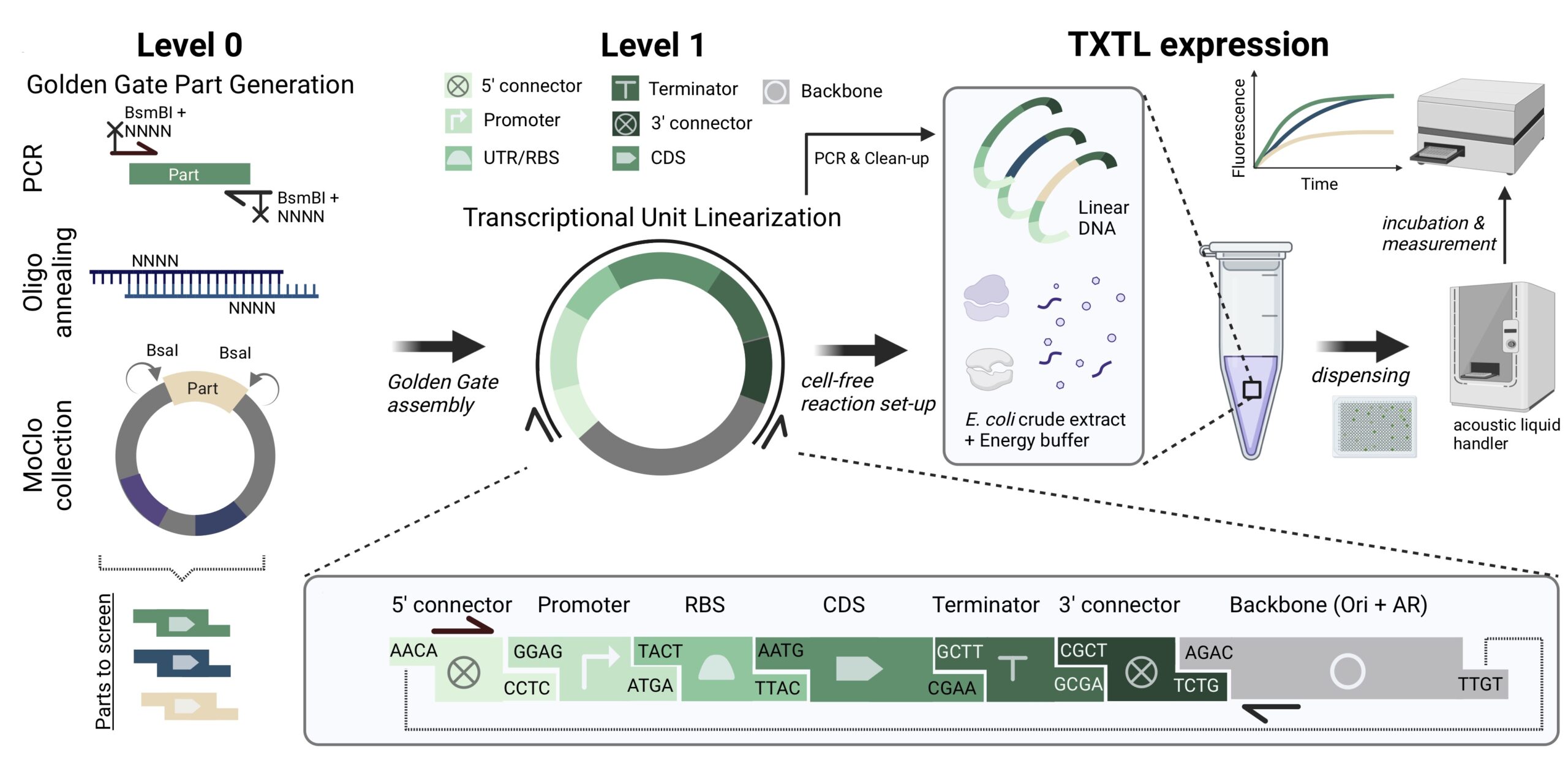
Modular Golden Gate Assembly of Linear DNA Templates for Cell-free Prototyping.
Lehr FX*, Gaizauskaite A*, Lipinska KE, Gilles S, Sahoo A, Inckemann R, Niederholtmeyer H#
In: Schindler, D. (eds) Golden Gate Cloning. Methods in Molecular Biology, vol. 2850. (2024)
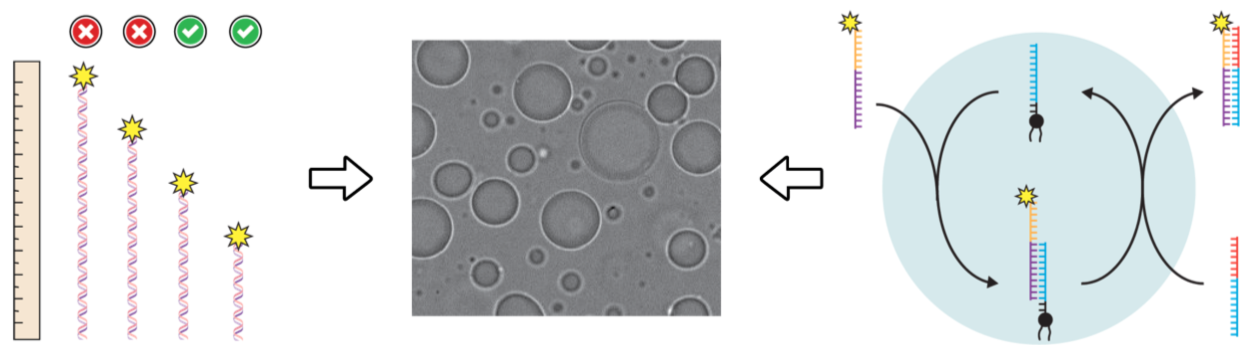

Functionalizing lipid sponge droplets with DNA.
Cho C, Niederholtmeyer H#, Seo H, Bhattacharya A, Devaraj NK#
ChemSystemsChem 10.1002/syst.202100045. (2022)
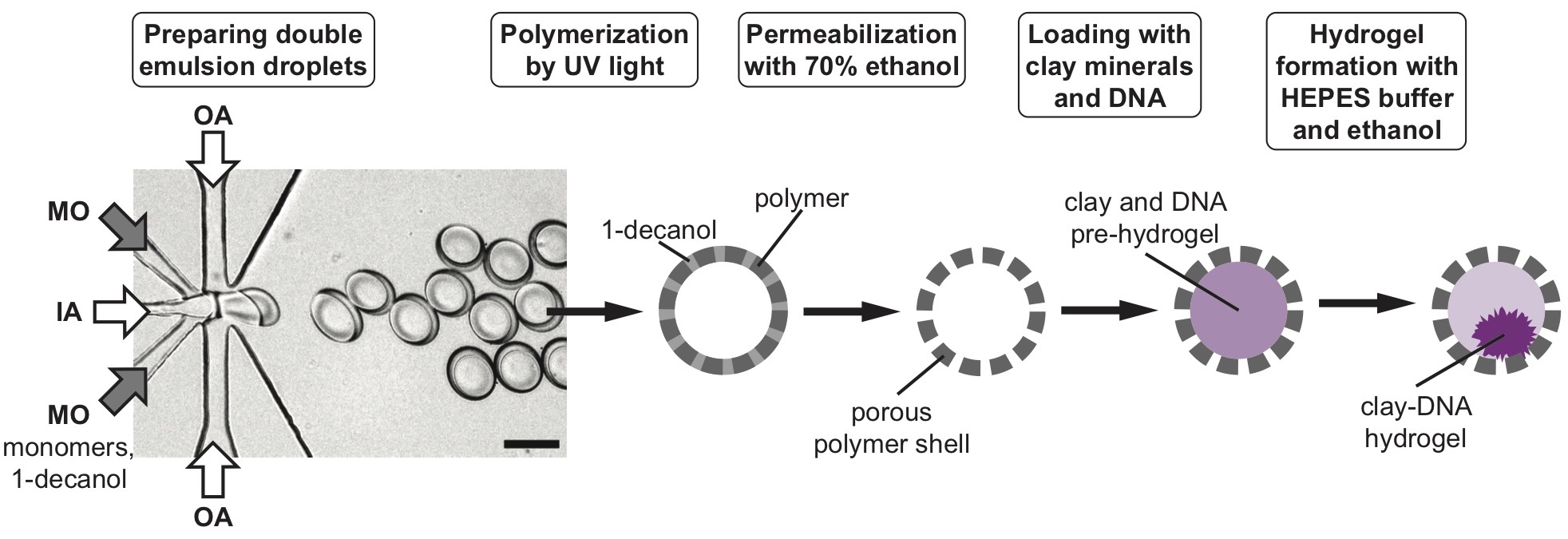
Microfluidic production of porous polymer cell-mimics capable of gene expression.
Banlaki I, Lehr FX, Niederholtmeyer H#
In: Karim, A.S., Jewett, M.C. (eds) Cell-Free Gene Expression. Methods in Molecular Biology, vol. 2433. (2022)
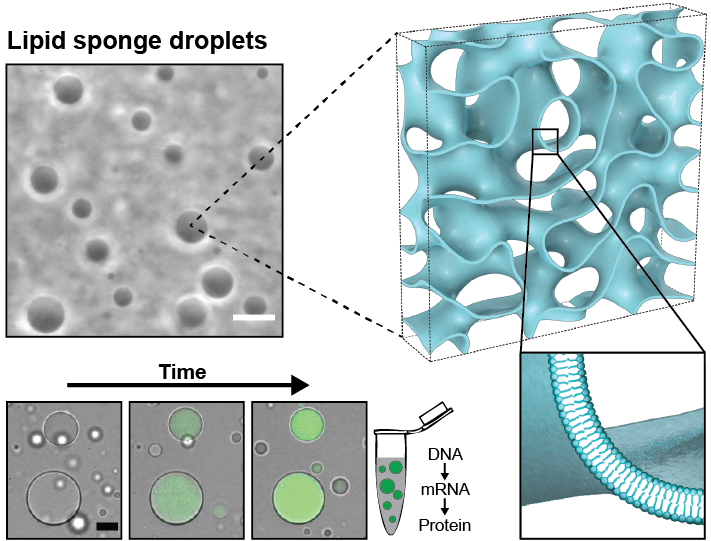
Lipid sponge droplets as programmable synthetic organelles.
Bhattacharya A*, Niederholtmeyer H*, Podolsky KA, Bhattacharya R, Song JJ, Brea RJ, Tsai CH, Sinha SK, Devaraj NK
Proceedings of the National Academy of Sciences 10.1073/pnas.2004408117. (2020)
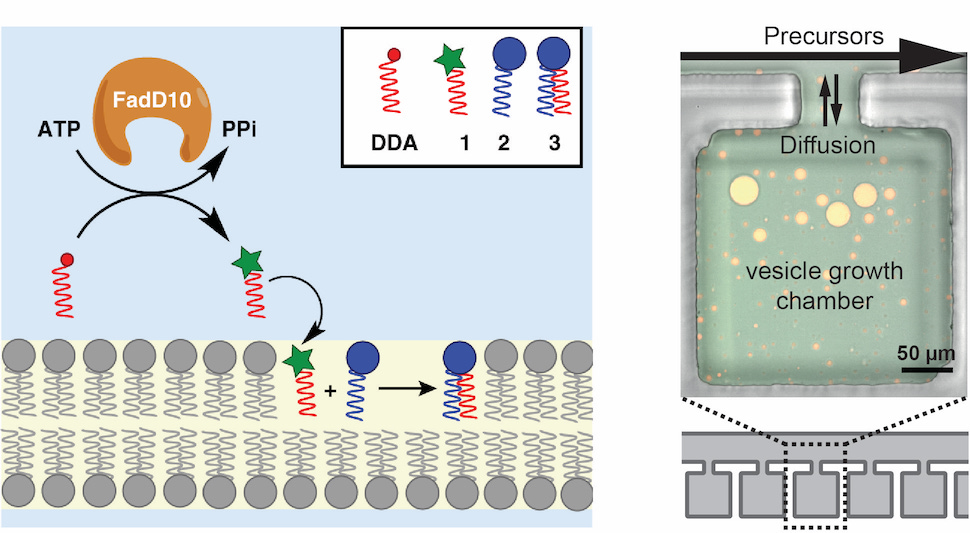
A minimal biochemical route towards de novo formation of synthetic phospholipid membranes.
Bhattacharya A, Brea RJ, Niederholtmeyer H, Devaraj NK
Nature Communications 10, 300. (2019)
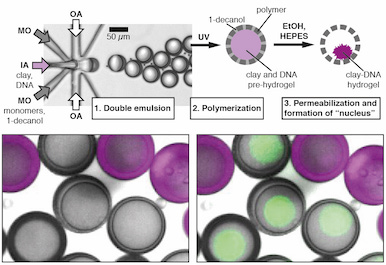
Communication and quorum sensing in non-living mimics of eukaryotic cells.
Niederholtmeyer H, Chaggan C, Devaraj NK
Nature Communications 9, 5027. (2018)
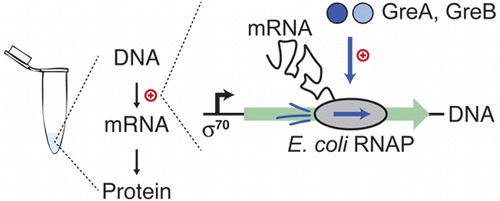
GreA and GreB enhance expression of Escherichia coli RNA polymerase promoters in a reconstituted transcription-translation system.
De Maddalena LL*, Niederholtmeyer H*, Turtola M, Swank Z, Belogurov GA, Maerkl SJ
ACS Synthetic Biology 5, 929-935. (2016)
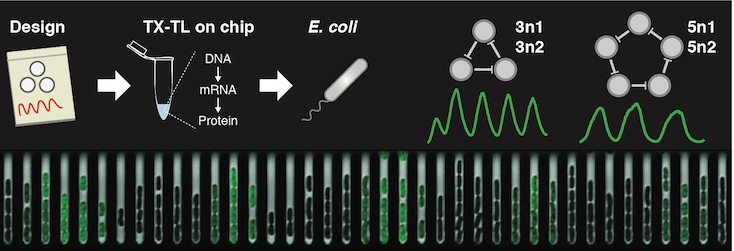
Rapid cell-free forward engineering of novel genetic ring oscillators.
Niederholtmeyer H*, Sun ZZ*, Hori Y, Yeung E, Verpoorte A, Murray RM, and Maerkl SJ
eLife 4, e09771. (2015).
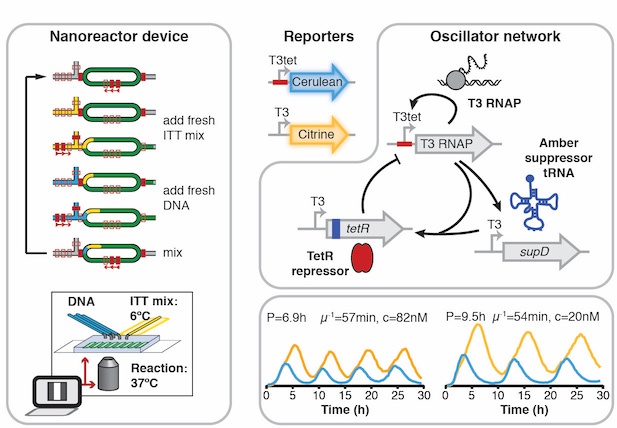
Implementation of cell-free biological networks at steady state.
Niederholtmeyer H, Stepanova V, and Maerkl SJ
Proceedings of the National Academy of Sciences 110, 15985–15990. (2013)
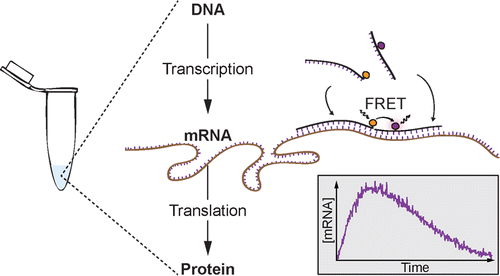
Real-time mRNA Measurement during an in vitro transcription and translation reaction using binary probes.
Niederholtmeyer H, Xu L, and Maerkl SJ
ACS Synthetic Biology 2, 411–417. (2012)
Modularity of a carbon-fixing protein organelle.
Bonacci W, Teng PK, Afonso B, Niederholtmeyer H, Grob P, Silver PA, and Savage DF
Proceedings of the National Academy of Sciences 109, 478–483. (2012)
Towards a synthetic chloroplast.
Agapakis CM, Niederholtmeyer H, Noche RR, Lieberman TD, Megason SG, Way JC, and Silver PA
PLoS ONE 6, e18877. (2011)
Engineering cyanobacteria to synthesize and export hydrophilic products.
Niederholtmeyer H*, Wolfstädter BT*, Savage DF, Silver PA, and Way JC
Applied and Environmental Microbiology 76, 3462. (2010)
Polyphosphate/ATP-dependent NAD kinase of Corynebacterium glutamicum: biochemical properties and impact of ppnK overexpression on lysine production.
Lindner SN*, Niederholtmeyer H*, Schmitz K, Schoberth SM, and Wendisch VF
Applied Microbiology and Biotechnology 87, 583–593. (2010)
* equal contribution
# corresponding author